Learning on Arbitrary Graph Topologies via Predictive Coding
To solve distinct learning tasks, artificial neural networks typically need to have different architectures and be trained in different ways. This paper shows that a model of information processing in the brain can solve all this tasks within a single network.
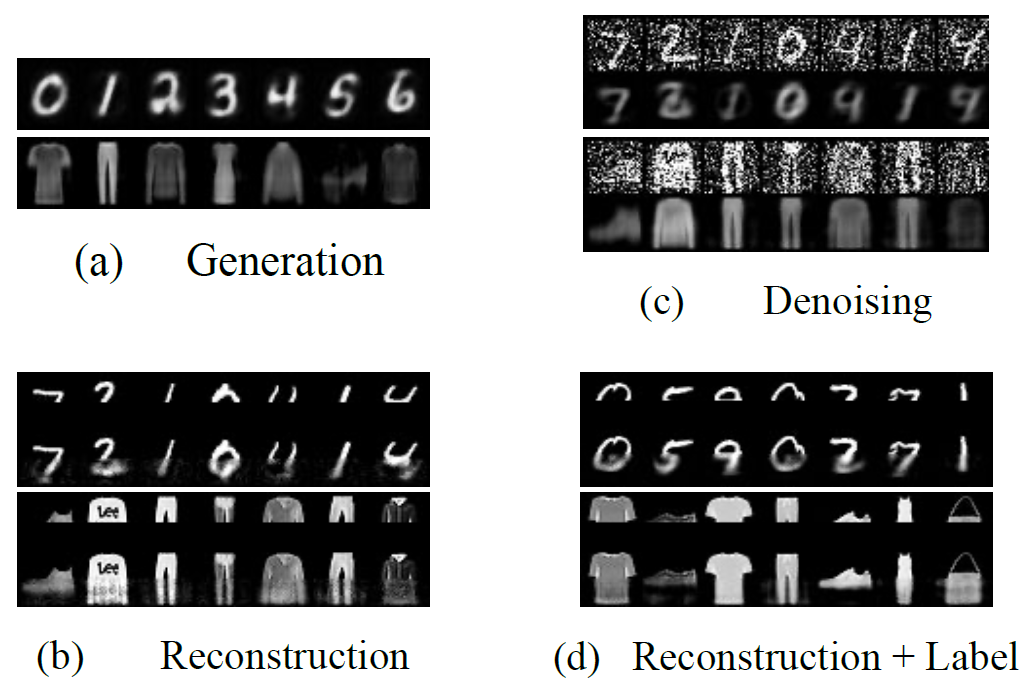
Training with backpropagation (BP) in standard deep learning consists of two main steps: a forward pass that maps a data point to its prediction, and a backward pass that propagates the error of this prediction back through the network. This process is highly effective when the goal is to minimize a specific objective function. However, it does not allow training on networks with cyclic or backward connections. This is an obstacle to reaching brain-like capabilities, as the highly complex heterarchical structure of the neural connections in the neocortex are potentially fundamental for its effectiveness. In this paper, we show how predictive coding (PC), a theory of information processing in the cortex, can be used to perform inference and learning on arbitrary graph topologies. We experimentally show how this formulation, called PC graphs, can be used to flexibly perform different tasks with the same network by simply stimulating specific neurons. This enables the model to be queried on stimuli with different structures, such as partial images, images with labels, or images without labels. We conclude by investigating how the topology of
the graph influences the final performance, and comparing against simple baselines trained with BP.